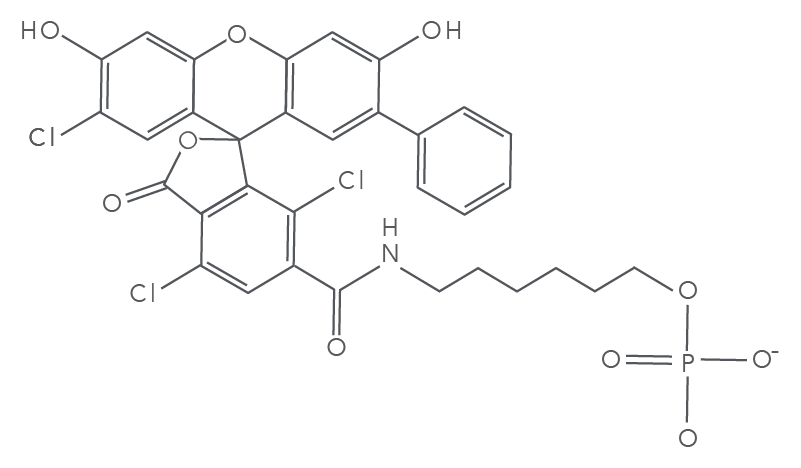
SUN fluorophore characteristics
Fluorophore characteristics such as absorption and emission wavelengths and intensity of the emitted light are key to providing clear and accurate gene expression and genotyping experimental results. SUN fluorophore has peak excitation at 538 nm and peak emission at 554 nm, spectral characteristics that are similar to HEX and VIC dyes. In addition, most common qPCR instruments can read the emission without additional calibration. SUN dye is a new addition to the portfolio of Freedom™ Dyes available from IDT, providing you with a license-free alternative to VIC dye. See Freedom Dyes for more information.
Table 1. Common fluorophore emission and excitation wavelengths.
| Fluorescent dye(s) | Excitation wavelength (nm) |
Emission wavelength (nm) |
|---|---|---|
| FAM | 495 | 520 |
| SUN | 538 | 554 |
| VIC | 538 | 554 |
| HEX | 538 | 555 |
| Cy® 3 | 550 | 564 |
| Texas Red® -X | 598 | 617 |
| Cy® 5 | 648 | 667 |
VIC is a trademark of Applied BioSystems;Yakima Yellow is a registered trademark of ELITechGroup; Texas Red-X is licensed from Molecular Probes, Inc., a wholly owned subsidiary of Life Technologies, Corp.; Cy is a registered trademark of Cytiva.
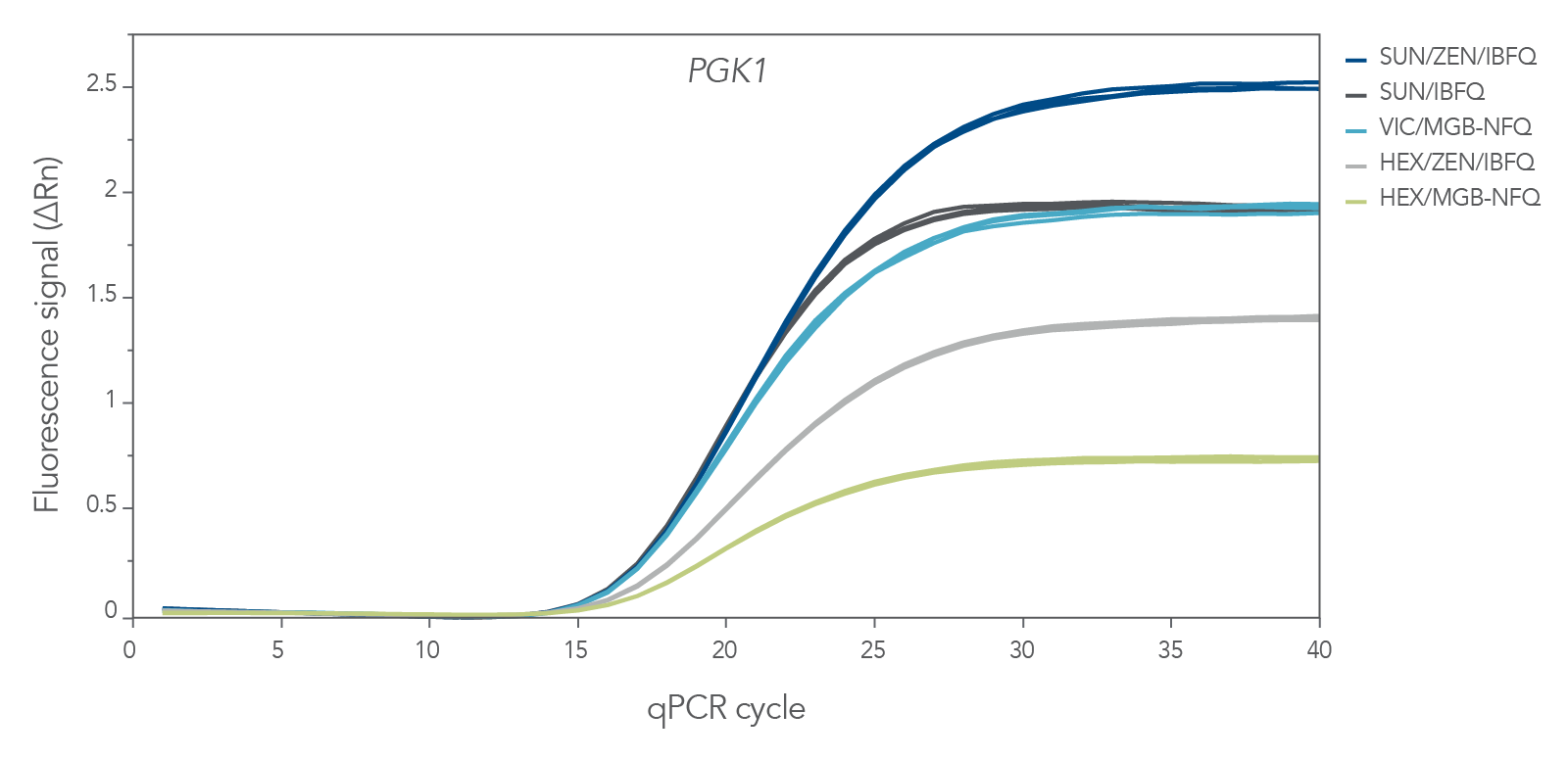
SUN probes provide strong signal intensity
Applications such as qPCR, multiplex qPCR, and digital PCR require oligonucleotide probes with strong emission spectra, especially for rare transcript detection. In qPCR assays containing primers specific to PGK1, the fluorescence signal for SUN probes after background subtraction (ΔRn) was comparable to or higher than the probes labeled with VIC or HEX dyes.
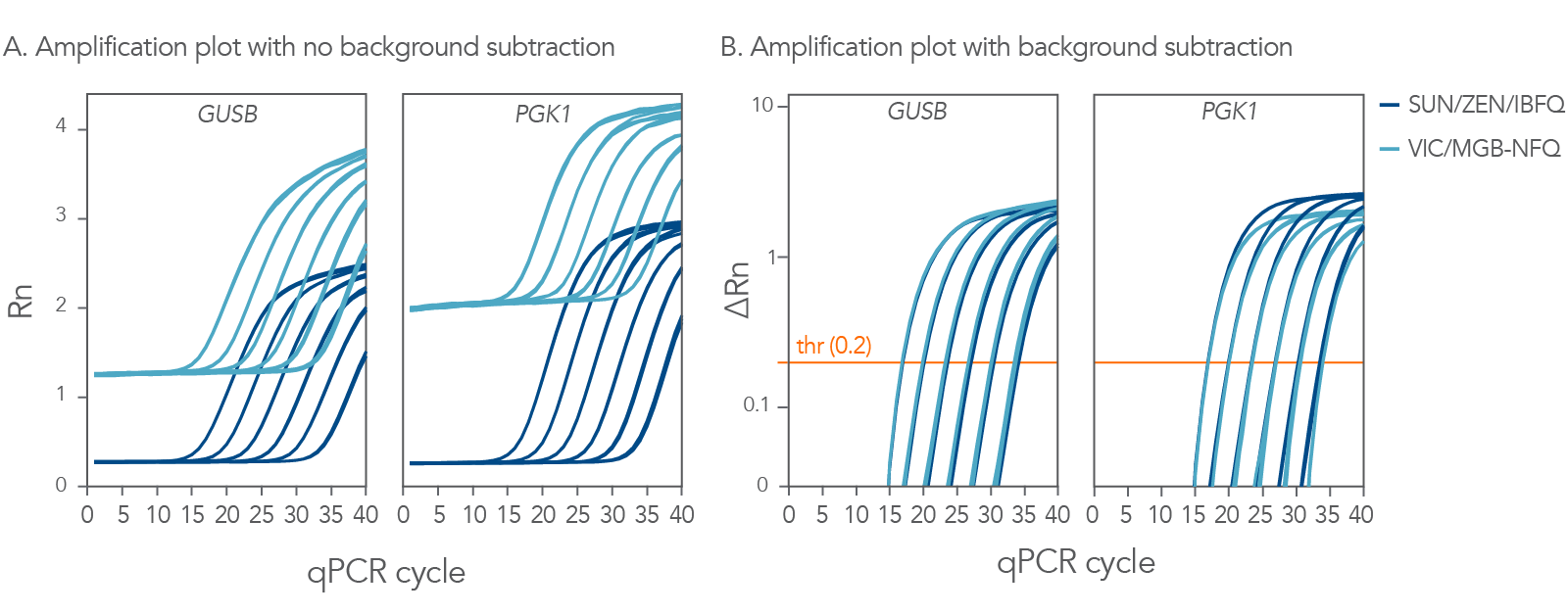
SUN probes perform comparable to VIC probes in qPCR
SUN fluorophore is a molecular equivalent to VIC dye and provides comparable or better signal intensity compared to dyes with similar excitation and emission spectra. To investigate SUN fluorophore performance, qPCR was performed on 2 different target genes, GUSB and PGK1. The assays used identical primers, but to directly compare 5’ nuclease probes, one was labeled with a 5’ SUN dye and the other was labeled with a 5’ VIC fluorophore. The SUN-labeled probe had excellent signal intensity, performing comparably to VIC-labeled probes for both gene targets. In part A, the lower starting fluorescence value of the SUN labeled probe is primarily due to quencher configuration. When the background levels of fluorescence were subtracted (Figure 3, part B), the Cq values were equivalent, and the resulting standard curves for both probes had R2 values of 0.999.
These data clearly demonstrate that the 5’ SUN-labeled probe performs comparably in gene expression studies and can outperform VIC-labeled probes in fluorescent intensity.
SUN probes are available in many different configurations
SUN probes can be purchased as individual PrimeTime qPCR probes or as part of PrimeTime qPCR Probe Assays. They are available as both double-quenched and single-quenched probes in a variety of different sizes and scales for gene expression studies. The probes can be ordered with mixed bases or locked nucleic acid bases, such as Affinity Plus monomers. Locked nucleic acids significantly increase Tm for shorter probe designs. Modulating the number of Affinity Plus bases in each probe for genotyping applications. See Affinity Plus qPCR Probes for more information.
5’ SUN-labeled nuclease probes are a license-free option for obtaining high-quality qPCR data from your gene expression and genotyping applications.